Intervertebral disc degeneration is accompanied by lysosomal impairment
Since lysosomal dysfunction represents one of many important contributors to illness development, we sought to research the connection between lysosomal perform and IDD. We categorized and analyzed the degenerative grades of human intervertebral disc samples in keeping with the MRI photographs. Primarily based on the histological staining (Fig. 1A), samples with larger degenerative grades, corresponding to grades IV and V, exhibited extra densely organized tissue buildings, elevated blue depth in Masson staining, and elevated inexperienced areas in Safranin O-fast inexperienced staining. These findings collectively recommend collagen loss and elevated fibrotic adjustments in degenerated intervertebral discs (Fig. 1A). Galectin-3 serves as a marker for lysosomal impairment, and we noticed that its protein ranges progressively elevated with larger grades of disc degeneration (Fig. 1B), alongside elevated expression of the LGALS3 gene in human NP tissues (Fig. 1C). Correlation evaluation revealed a optimistic linear relationship between LGALS3 gene expression ranges and IDD grades (Fig. 1D). Moreover, immunofluorescence co-localization of galectin-3 with the lysosomal protein LAMP1 demonstrated progressively growing galectin-3 expression inside lysosomes as disc degeneration superior (Fig. 1E).
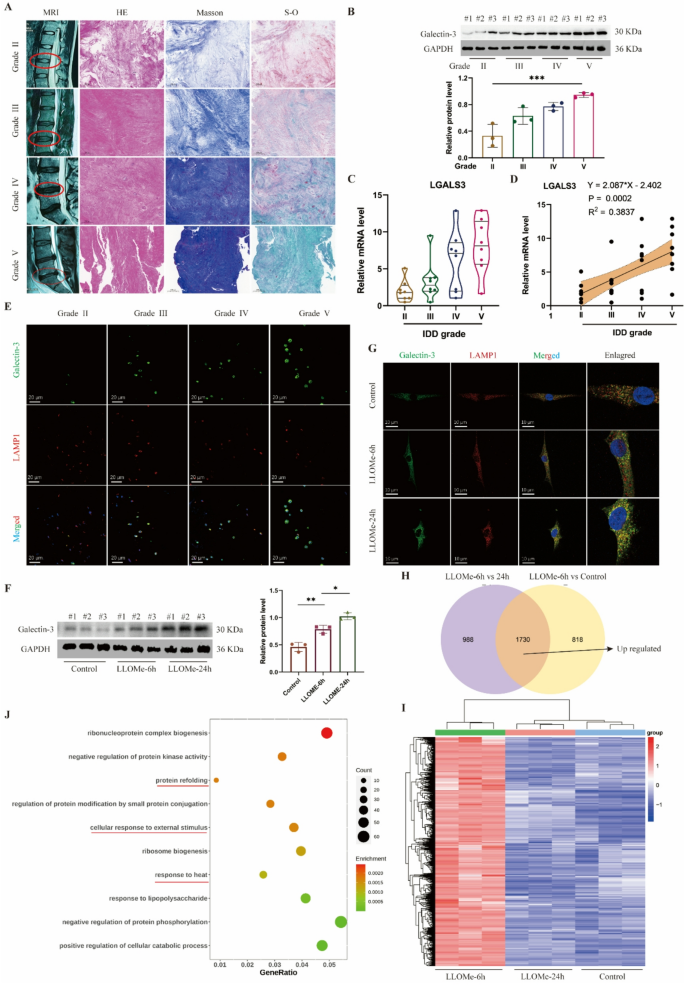
Lysosomal dysfunction is related to IDD and NP cell harm. A, Consultant MRI photographs, HE, Masson and Safranin O-fast inexperienced (S-O) staining of human intervertebral disc samples with totally different degenerative grades (Grade II, III, IV and V). B, Western blot of Galectin-3 and quantitative evaluation of protein ranges in human intervertebral disc samples. C, Relative mRNA stage of LGALS3 gene in human intervertebral disc samples. D, Correlation evaluation between the MRI grade and the expression stage of LGALS3. E, Consultant immunofluorescence of Galectin-3 and LAMP1 in human intervertebral disc samples. F, Western blot of Galectin-3 and quantitative evaluation of protein ranges in human NP cells handled with LLOMe (100 μm) at 0, 6 and 24 h. G, Consultant immunofluorescence of Galectin-3 and LAMP1 in human NP cells handled with LLOMe. H, Venn plot of up-regulated expressed genes in LLOMe-treated 6 h group. I, Heatmap of up-regulated expressed genes in LLOMe-treated 6 h group. J, GO enrichment of up-regulated expressed genes in LLOMe-treated 6 h group. Knowledge are proven because the imply ± SD (n ≥ 3). *p < 0.05, **p < 0.01, ***p < 0.001 by a technique ANOVA, or linear regression
To induce lysosomal impairment on the mobile stage, we handled human NP cells with LLOMe, a generally used lysosomal injury inducer. As LLOMe remedy length elevated, galectin-3 protein expression ranges in NP cells confirmed a progressive rise (Fig. 1F), whereas immunofluorescence outcomes confirmed elevated galectin-3 localization inside lysosomes (Fig. 1G). Concurrent with lysosomal harm, LLOMe remedy induced NP cell apoptosis, with essentially the most pronounced impact noticed on the 24-hour remedy time level (Determine S1A). To determine key regulatory molecules concerned in lysosomal restore inside NP cells, we carried out transcriptome sequencing on human NP cells handled with LLOMe for 0, 6, and 24 h (Determine S1B). Our evaluation centered on genes upregulated on the 6-hour timepoint, as these may comprise crucial early-response molecules for lysosomal harm restore. In comparison with management and 24-hour LLOMe-treated teams (Fig. 1H-I), the 6-hour remedy group confirmed 1,730 differentially upregulated genes (p-value < 0.05). GO enrichment evaluation revealed these genes have been primarily related to “protein refolding” and responses to “warmth or stimulus” (Fig. 1J), suggesting mobile self-regulative mechanisms throughout acute harm phases. KEGG pathway evaluation indicated that the upregulated genes have been associated to inflammatory signaling pathways together with TNF-α, NF-κB, and IL-17 signaling (Determine S1C). REACTOME evaluation additional demonstrated connections to mobile warmth shock responses and inflammatory reactions (Determine S1D). Collectively, our findings exhibit a robust affiliation between lysosomal injury and IDD. Nonetheless, the precise molecules collaborating in lysosomal restore processes in NP cells require additional elucidation.
RRAGD serves as a key molecule for sustaining lysosomal perform
From the pool of 1,730 differentially upregulated genes, we screened these genes with perform that associated to “lysosome” and “stress response.” Amongst these, 4 genes, together with RRAGC, RRAGD, MIOS and RNF152, met the choice standards. We subsequently established a number of lysosomal harm fashions utilizing TBHP, LLOMe, and silica to induce NP cell injury. Outcomes confirmed that RRAGD expression ranges have been considerably elevated in handled teams throughout all cell fashions in comparison with controls (Determine S2A). For additional validation, we examined protein and RNA ranges of RRAGD in human NP tissues at totally different degenerative grades. Our evaluation revealed progressively lowering RRAGD protein (Fig. 2A) and mRNA expression stage (Fig. 2B) as disc degeneration superior. Correlation evaluation confirmed a damaging linear relationship between RRAGD expression ranges and IDD degenerative grades (Fig. 2C). Notably, whereas the 6-hour LLOMe remedy group confirmed no important change in RRAGD protein ranges in comparison with controls, a considerable lower occurred after 24 h of remedy (Determine S2B). Conversely, RRAGD transcriptional ranges have been considerably elevated after 6 h of LLOMe publicity however decreased in LLOMe handled 24-hour group (Determine S2C). Immunofluorescence outcomes demonstrated RRAGD co-localization with LAMP1 on the 6-hour time level, which turned much less evident after 24 h (Fig. 2D). These findings recommend that whereas RRAGD gene transcription will increase considerably throughout early-stage stress, its expression markedly declines throughout later phases of harm.
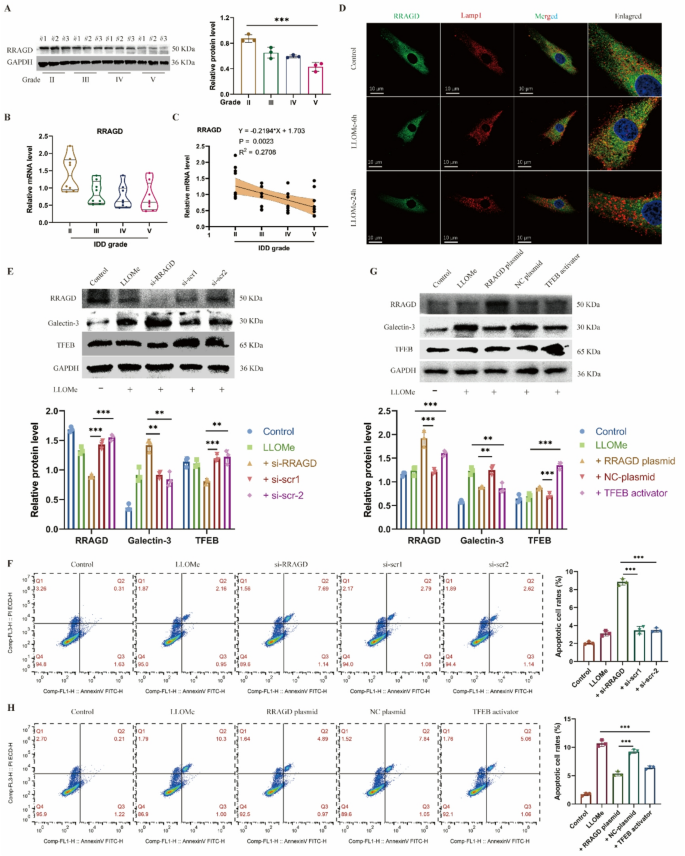
RRAGD maintains the lysosomal perform in human NP cells. A, Western blot of RRAGD and quantitative evaluation of protein ranges in human intervertebral disc samples with totally different degenerative grades (Grade II, III, IV and V). B, Relative mRNA stage of RRAGD gene in human intervertebral disc samples. C, Correlation evaluation between the MRI grade and the expression stage of RRAGD. D, Consultant immunofluorescence of RRAGD and LAMP1 in human NP cells handled with LLOMe. E, Western blot of RRAGD, Galectin-3 and TFEB and quantitative evaluation of protein ranges in human NP cells transfected with RRAGD siRNA or scrambled siRNA. F, Movement cytometry photographs of Annexin V/PI and quantification of apoptotic cell charge in respective teams. G, Western blot of RRAGD, Galectin-3 and TFEB and quantitative evaluation of protein ranges in human NP cells transfected with RRAGD plasmid or NC plasmid or handled with TFEB activator. H, Movement cytometry photographs of Annexin V/PI and quantification of apoptotic cell charge in respective teams. Knowledge are proven because the imply ± SD (n ≥ 3). *p < 0.05, **p < 0.01, ***p < 0.001 by pupil’s t check, a technique ANOVA, or linear regression
To additional examine RRAGD perform, we employed siRNA to inhibit the expression of RRAGD. Following remedy of RRAGD-knockdown NP cells with LLOMe, galectin-3 confirmed elevated expression whereas TFEB displayed lowered ranges, indicating the compromised lysosomal biogenesis (Fig. 2E). Cell viability assays indicated considerably decreased NP cell exercise after RRAGD interference (Determine S2D). Furthermore, RRAGD knockdown exacerbated apoptosis in LLOMe-treated NP cell (Fig. 2F). Conversely, transfection with RRAGD-overexpressing plasmids elevated each RRAGD and TFEB expression in NP cells (Fig. 2G). The RRAGD overexpression group exhibited considerably a decrease stage of galectin-3 in comparison with overexpression management, with results akin to TFEB activator remedy (Fig. 2G). Notably, RRAGD overexpression promoted restoration of cell viability (Determine S2F) and lowered cell apoptosis in comparison with LLOMe remedy alone (Fig. 2H). In abstract, RRAGD performs a vital position in sustaining regular lysosomal perform, whereas its deficiency exacerbates lysosomal injury in IDD.
The Fc-TRIM21 system supplies an environment friendly platform for EV protein loading
The PRYSPRY area of the Fc fragment demonstrates sturdy binding to its cytoplasmic receptor TRIM21. We deliberate to make the most of this Fc-TRIM21 interplay to realize focused RRAGD loading and transport inside EVs, finally aiming for lysosomal injury restore. Initially, we evaluated a number of candidate EV membrane scaffold proteins together with PTGFRN, PDGFR, LAMP2B, CD9, and CD63 to determine essentially the most appropriate platform (Fig. 3A). Co-transfection of EV scaffold protein and RRAGD plasmids into 293T cells enabled the manufacturing of engineered EVs (Fig. 3B). The outcomes confirmed detectable RRAGD in all scaffold-based engineered EVs, although expression ranges diversified (Fig. 3C). All engineered EV preparations have been damaging for the endoplasmic reticulum protein calnexin, indicating the purity of EVs. Coomassie blue staining revealed typically comparable protein composition patterns throughout several types of engineered EV (Fig. 3C). When equal quantities of engineered EVs have been administered to NP cells, the PTGFRN group confirmed the very best exogenous RRAGD stage (Determine S3A). Utilizing fluorescent His-tag to trace RRAGD-containing EV uptake, we noticed inexperienced fluorescent indicators in all engineered EV remedy teams, with the PTGFRN group displaying the strongest depth (Determine S3B). The stream cytometry evaluation confirmed that over 95% of NP cells internalized the engineered EVs. In comparison with controls, the engineered EV-treated group confirmed considerably elevated imply fluorescent depth (MFI), with PTGFRN-EVs demonstrating the very best MFI (Fig. 3D-E). Primarily based on these outcomes, PTGFRN was chosen because the optimum scaffold protein for EV-based supply system.
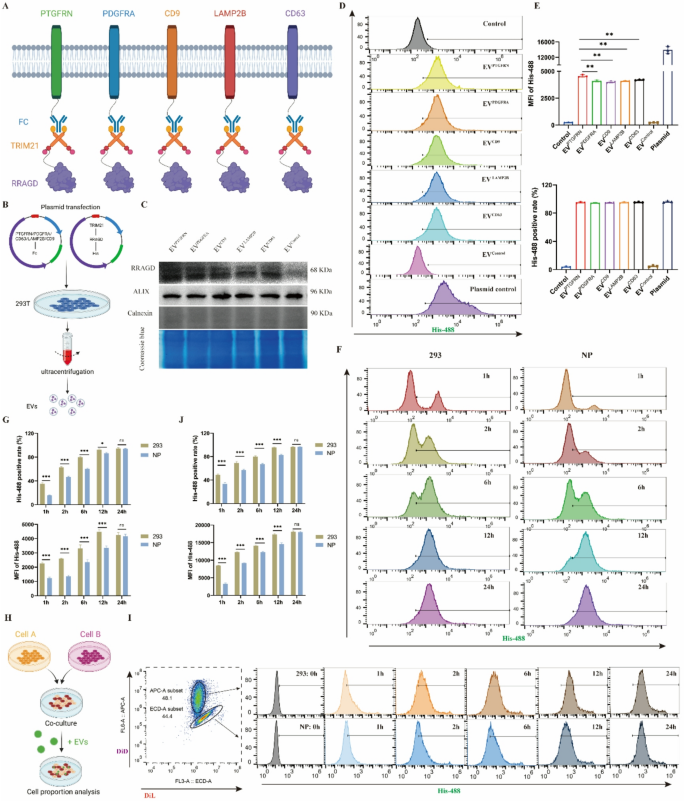
Building of EV-based protein loading system. A, Schematic diagram of Fc/TRIM21 system to ship RRAGD protein based mostly on totally different EV scaffold proteins. B, Workflow of isolating engineered EVs. C, Western blot of RRAGD, Alix and Calnexin in respective engineered EVs and Coomassie blue staining. D, Movement cytometry photographs of His-488-labeled engineered EVs incubated with human NP cells. E, Imply fluorescent depth (MFI) of His-488 and His-488 optimistic cell charge in respective teams. F, Movement cytometry photographs of His-488-labeled PTGFRN-engineered EVs incubated with human NP cells or 293T cells at 1, 2, 6, 12, and 24 h. G, MFI of His-488 and His-488 optimistic cell charge in respective teams. H, Workflow of cell direct tradition mannequin development and cell labeling. I, Movement cytometry photographs of His-488-labeled PTGFRN-engineered EVs incubated with DiL-labeled NP cells or DID-labeled 293T cells at 0, 1, 2, 6, 12, and 24 h. J, MFI of His-488 and His-488 optimistic cell charge in respective teams. Knowledge are proven because the imply ± SD (n ≥ 3). *p < 0.05, **p < 0.01, ***p < 0.001, ns (not important) by a technique or two means ANOVA
Utilizing PTGFRN because the scaffold protein and leveraging the Fc-TRIM21 binding platform, we constructed RRAGD-delivering engineered EVs. We additional investigated potential variations in EV uptake effectivity between totally different cell varieties. Equal quantities of engineered EVs have been administered to human NP cells and 293T cells, and EV uptake was monitored. Outcomes confirmed growing proportions of EV-positive cells over time (Fig. 3F-G), although NP cells reached the height proportion extra slowly than 293T cells. To get rid of potential tradition environmental results on uptake effectivity, we established a direct coculture mannequin with NP cells and 293T cells at a 1:1 ratio (Fig. 3H). Utilizing totally different fluorescent dyes to label NP cells and 293T cells, we analyzed intracellular fluorescence depth adjustments following engineered EV remedy at varied time factors. Each NP cells and 293T cells confirmed growing MFI and EV-positive cell proportions over time. Nonetheless, 293T cells achieved each most EV-positive cell share and highest MFI sooner than NP cells (Fig. 3I-J). These findings point out that NP cells exhibit decrease EV uptake effectivity in comparison with 293T cells, which can influence therapeutic effectiveness of engineered EVs in NP cells.
CAP-engineered EVs possess NP cell-targeting functionality
To enhance EV uptake effectivity in human NP cells, we optimized the EV membrane floor to boost cell concentrating on. CAP peptide, a cartilage-affinity peptide with superb binding to NP cells, was fused to the extracellular area of PTGFRN (Fig. 4A), creating an engineered EVs (EVPTGFRN−CAP). The outcomes confirmed that CAP peptide expression didn’t have an effect on bodily properties of engineered EVs, together with their morphology and dimension distribution (Determine S4A). Comparability between EVPTGFRN and EVPTGFRN−CAP revealed no important variations in RRAGD and PTGFRN protein ranges, indicating that CAP modification makes no impact on EV protein loading. The Coomassie blue staining additionally indicated minimal influence of CAP peptide incorporation on EV protein composition (Fig. 4B). When handled with equal quantities of EVPTGFRN−CAP and EVPTGFRN for an identical durations, the NP cells in EVPTGFRN−CAP group confirmed considerably larger stage of exogenous RRAGD (Determine S4B). To additional analyze the variations of EV uptake effectivity, we used fluorescent His-tagged RRAGD to trace EV internalization in human NP cells. Each EVPTGFRN−CAP and EVPTGFRN teams may detect inexperienced fluorescence indicators, with larger MFI within the CAP-modified EV group (Determine S4C). The stream cytometry outcomes confirmed comparable proportions of EV-positive cells between two teams after 24-hour EV remedy, however EVPTGFRN−CAP exhibited larger MFI of His-488 sign (Fig. 4C-D). These outcomes point out that NP cells exhibit enhanced uptake effectivity for CAP-optimized engineered EVs.
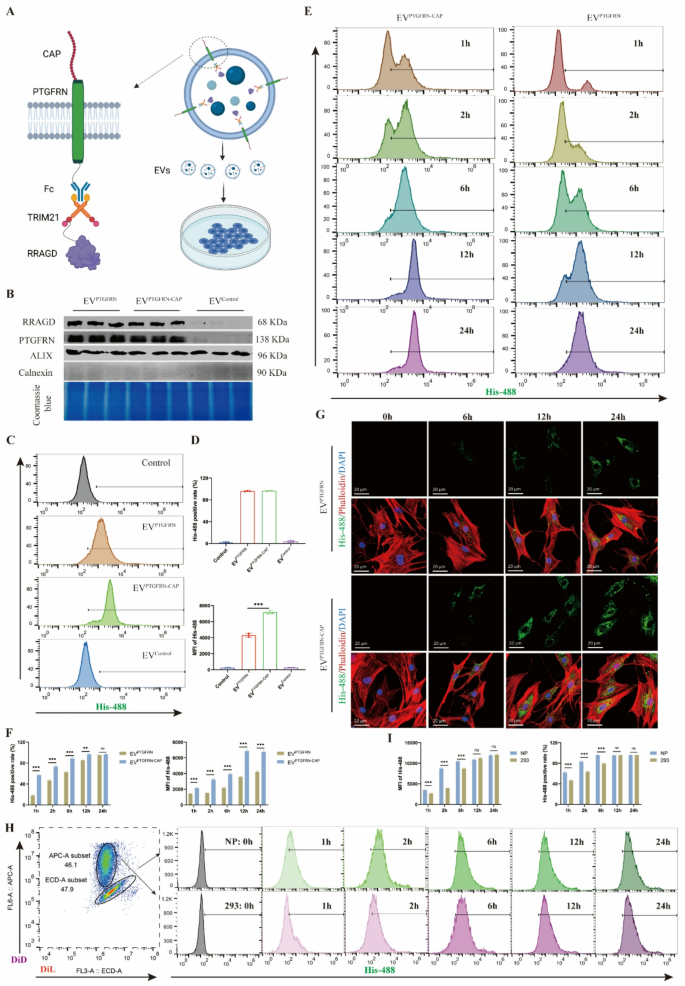
Engineering EVs with CAP peptide to acquire NP cell concentrating on. A, Schematic diagram of CAP-modified engineered EVs based mostly on Fc/TRIM21 system. B, Western blot of RRAGD, PTGFRN, Alix and Calnexin in respective engineered EVs and Coomassie blue staining. C, Movement cytometry photographs of His-488-labeled engineered EVs incubated with human NP cells. D, MFI of His-488 and His-488 optimistic cell charge in respective teams. E, Movement cytometry photographs of His-488-labeled EVPTGFRN−CAP or EVPTGFRN incubated with human NP cells at 1, 2, 6, 12, and 24 h. F, MFI of His-488 and His-488 optimistic cell charge in two teams. G, Consultant immunofluorescent photographs of His-488-labeled EVPTGFRN−CAP or EVPTGFRN incubated with human NP cells. Phalloidin for actin and DAPI for nucleus. H, Movement cytometry photographs of His-488-labeled EVPTGFRN−CAP incubated with DiL-labeled NP cells or DID-labeled 293T cells at 0, 1, 2, 6, 12, and 24 h. I, MFI of His-488 and His-488 optimistic cell charge in respective teams. Knowledge are proven because the imply ± SD (n ≥ 3). **p < 0.01, ***p < 0.001, ns (not important) by a technique or two means ANOVA
Prolonged time-course experiments revealed growing proportions of EV-positive cells and MFI over time within the EVPTGFRN−CAP and EVPTGFRN group. Nonetheless, the EVPTGFRN−CAP group reached peak values sooner than EVPTGFRN and achieved larger most MFI (Fig. 4E-F). Immunofluorescence staining additional confirmed persistently larger MFI within the EVPTGFRN−CAP group throughout all time factors (Fig. 4G, S4D). Apart from, coculture experiments have been performed to check the uptake of EVPTGFRN−CAP by NP cell and 293T cell. These experiments demonstrated that each cell varieties exhibited elevated MFI and EV-positive cell proportions over time. Notably, NP cells achieved peak MFI quickly throughout coculture, reaching most EV-positive cell share and peak MFI sooner than 293T cells (Fig. 4H-I). Moreover, immunofluorescence demonstrated colocalization of internalized engineered EVs with lysosomal membrane protein LAMP1 (Determine S4E), suggesting the environment friendly supply of EVs to NP cell lysosomes. In conclusion, these findings exhibit that CAP peptide modification confers NP cell concentrating on functionality to engineered EVs and considerably improves the EV uptake effectivity in NP cells.
Therapeutic results of engineered EVs on NP cells in vitro
Primarily based on earlier outcomes, we constructed engineered EVs expressing CAP peptide and delivering RRAGD by way of the PTGFRN-Fc-TRIM21 platform, then evaluated the consequences in NP cell harm fashions. Within the LLOMe-induced harm mannequin, each endogenous and exogenous RRAGD expression have been detected within the engineered EV-treated group. In comparison with the LLOMe-treated group, the engineered EV group exhibited considerably decreased the galectin-3 expression and elevated the TFEB expression stage (Fig. 5A). Whereas LLOMe remedy induced NP cell apoptosis, engineered EV administration markedly inhibited this impact (Fig. 5B). Moreover, LLOMe remedy lowered each the common MFI of Calcein-staining dwell cells and total cell viability, whereas engineered EVs partially restored cell survival (Fig. 5C, S5A-B). In further experiments, we remoted NP cells from non-degenerated (N-NP) and degenerated (D-NP) intervertebral discs and incubated them with engineered EVs. The D-NP group displayed larger galectin-3 and decrease TFEB expression than N-NP cells. Engineered EV remedy lowered the galectin-3 stage in each teams (Fig. 5D). Beneath customary tradition circumstances with out exogenous stress, neither group confirmed important apoptosis (Fig. 5E) or variations in cell viability (Fig. 5F, S5C-D), confirming that engineered EVs didn’t have an effect on the baseline of cell viability. To simulate inflammatory harm, we handled NP cells with IL-1β, which markedly upregulated the galectin-3 stage and downregulated the TFEB expression. Engineered EVs partially counteracted these IL-1β-induced results (Fig. 5G). IL-1β remedy additionally triggered substantial apoptosis, whereas engineered EVs considerably mitigated this impact (Fig. 5H). Though IL-1β lowered live-cell fluorescence depth and viability, engineered EVs partially restored NP cell well being beneath inflammatory circumstances (Fig. 5I, S5E-F). Collectively, these findings exhibit that engineered EVs exert therapeutic results on injured NP cells by enhancing viability and lowering apoptosis.
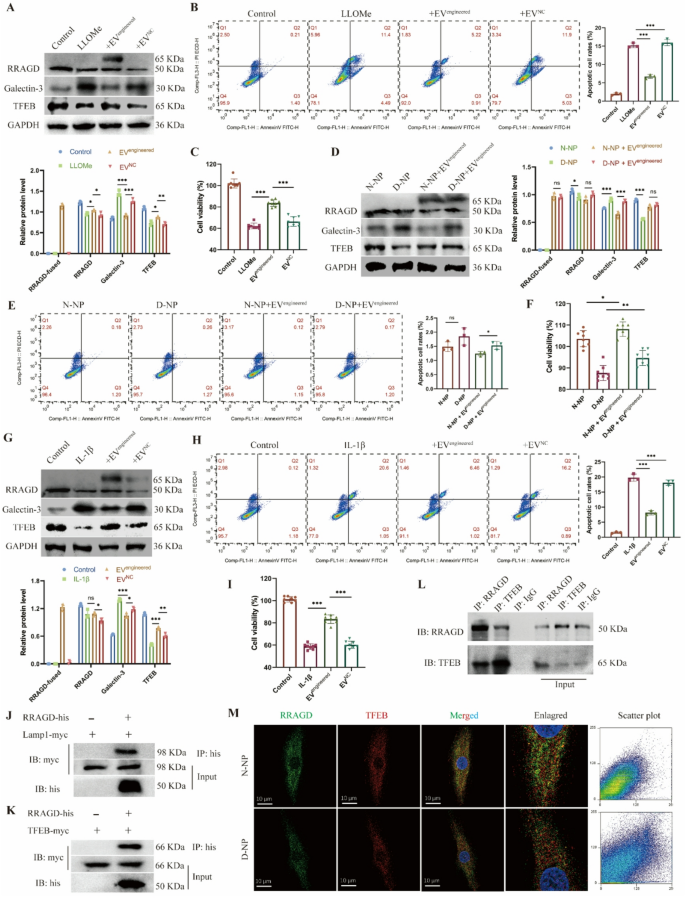
Therapeutic results of engineered EVs on human NP cells. A, Western blot of RRAGD, Galectin-3, TFEB, and quantitative evaluation of protein ranges in human NP cells handled with LLOMe and engineered or management EVs. B, Movement cytometry photographs of Annexin V/PI and quantification of apoptotic cell charge in respective teams. C, Cell viability assay in respective teams. D, Western blot of RRAGD, Galectin-3, TFEB, and quantitative evaluation of protein ranges in non-degenerated and degenerated human NP cells (N-NP and D-NP). E, Movement cytometry photographs of Annexin V/PI and quantification of apoptotic cell charge in respective teams. F, Cell viability assay in respective teams. G, Western blot of RRAGD, Galectin-3, TFEB, and quantitative evaluation of protein ranges in human NP cells handled with IL-1β and engineered or management EVs. H, Movement cytometry photographs of Annexin V/PI and quantification of apoptotic cell charge in respective teams. I, Cell viability assay in respective teams. J, Pull down assay of RRAGD-his and LAMP1-myc. Okay, Pull down assay of RRAGD-his and TFEB-myc. L, Co-immunoprecipitation assay of RRAGD and TFEB. IgG serves because the damaging management. M, Consultant immunofluorescent photographs and co-localization scatter plot of RRAGD and TFEB in N-NP and D-NP cells. Knowledge are proven because the imply ± SD (n ≥ 3). *p < 0.05, **p < 0.01, ***p < 0.001, ns (not important) by a technique ANOVA
To additional elucidate the therapeutic mechanism of engineered EVs, we analyzed the interactions of RRAGD with lysosomal proteins. We constructed plasmids expressing tagged RRAGD, LAMP1, and TFEB, which have been transfected into 293T cells additional. Pull-down assays confirmed a binding interplay between RRAGD and LAMP1 (Fig. 5J), suggesting the localization of RRAGD to lysosomal membranes. Moreover, our outcomes additionally revealed that RRAGD interacted with TFEB (Fig. 5K), suggesting its position in lysosomal biogenesis regulation. Co-immunoprecipitation in NP cells beneath physiological circumstances confirmed the binding relationship between RRAGD and TFEB (Fig. 5L). Immunofluorescence additional demonstrated RRAGD-TFEB co-localization in each non-degenerated and degenerated NP cells (Fig. 5M). These observations recommend that RRAGD localizes to lysosomes and features by means of binding with TFEB. In abstract, our engineered EVs successfully ship RRAGD to orchestrate lysosomal perform, thereby exerting therapeutic results on injured NP cells.
Therapeutic results of engineered EVs on disc degeneration in vivo
To systematically consider the therapeutic efficacy of engineered EVs in vivo, we delivered EVs into rat discs based mostly on a needle puncture-induced IDD mannequin. Disc degeneration development was comprehensively assessed by means of multimodal imaging (MRI, CT, X-ray) and histopathological evaluation. MRI quantified hydration adjustments in NP, whereas CT and X-ray detected structural alterations in intervertebral house (Fig. 6A). Degenerated discs exhibited considerably lowered hydration and collapsed intervertebral house in comparison with wholesome controls. The engineered EV group demonstrated decrease MRI grades and better Disc Peak Index (DHI) versus the IDD and EV management teams (Fig. 6B). Fluorescence imaging revealed comparable intradiscal accumulation between engineered and management EVs (Fig. 6C-D), whereas solely engineered EVs introduced results on IDD mitigation. Histological outcomes recognized degenerative traits of discs: NP tissue atrophy, annulus fibrosus (AF) disorganization with fiber rupture, and decreased disc peak with cartilaginous endplate (CEP) degeneration. The outcomes indicated that engineered EV considerably improved histological scores in comparison with the IDD and management EV teams (Fig. 6E-F).
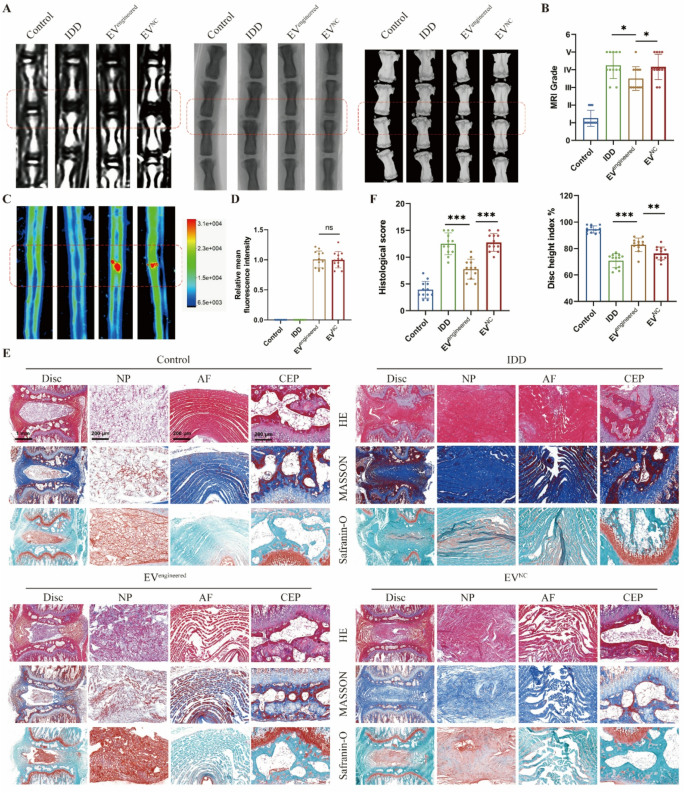
Engineered EVs retard the disc degeneration in vivo. A, MRI, X-ray and CT photographs of rat discs in respective teams. The rat disc was injected with engineered EVs (EVengineered) or management EVs (EVNC). B, MRI grades and Disc peak index based mostly on MRI, X-ray and CT photographs. C-D, Fluorescent photographs and relative imply fluorescence depth of rat discs in respective teams. E, HE, Masson and Safranin-O staining photographs of rat discs in respective teams. F, Histological scores based mostly on histopathological staining of rat discs in respective teams. Knowledge are proven because the imply ± SD (n = 12). *p < 0.05, **p < 0.01, ***p < 0.001, ns (not important) by a technique ANOVA
Moreover, immunofluorescence evaluation of RRAGD expression in rat discs demonstrated lowered ranges throughout disc compartments in IDD group versus management group. It revealed that engineered EVs selectively upregulated RRAGD expression in NP areas (Fig. 7A-B). It additionally confirmed that there was considerably decreased expression of galectin-3 in NP tissue of engineered EV-treated discs versus IDD and management EV group, with no important variations in AF and CEP areas (Fig. 7C-D). It demonstrated the NP-specific remedy and focused supply of engineered EVs. TUNEL staining confirmed an elevated cell apoptosis within the IDD group versus management group, whereas engineered EVs considerably lowered the speed of cell apoptosis (Fig. 7E-F). Collectively, engineered EVs successfully attenuated disc degeneration by means of focused RRAGD supply, exhibiting marked therapeutical potential for IDD.
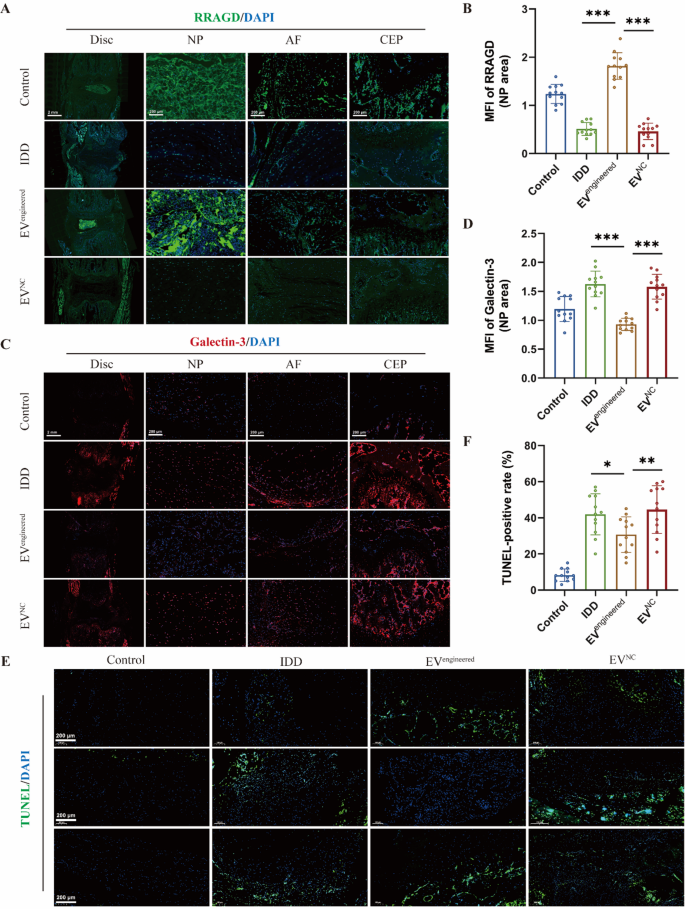
Engineered EVs ameliorate cell loss of life throughout disc degeneration in vivo. A, Immunofluorescence of RRAGD (inexperienced) in respective teams. B, Imply fluorescence depth of RRAGD within the rat NP space in respective teams. C, Immunofluorescence of galectin-3 (pink) in respective teams. D, Imply fluorescence depth of galectin-3 within the rat NP space in respective teams. E, TUNEL staining photographs of rat discs in respective teams. F, Quantification of TUNEL-positive cell charge in respective teams. Knowledge are proven because the imply ± SD (n = 12). *p < 0.05, **p < 0.01, ***p < 0.001 by a technique ANOVA

